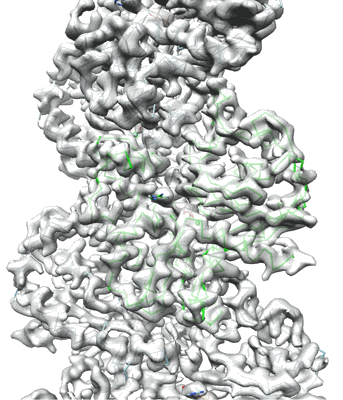
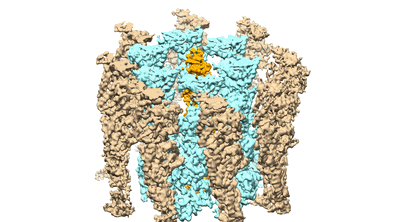
Resolution: 3.9 Å
EM Method: Helical reconstruction
Fitted PDBs: 7x54
Q-score: 0.375
Koh A, Ali S, Popp D, Tanaka K, Kitaoku Y, Miyazaki N, Iwasaki K, Mitsuoka K, Robert R, Narita A
To Be Published
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Cbg-parm filament with adp (Complex from Clostridium botulinum)
- Parm/stba family protein (32 kDa, Protein from Clostridium botulinum Bf)
- Magnesium ion (24 Da, Ligand)
A CBg-ParM filament with GTP and a short incubation time
Resolution: 8.6 Å
EM Method: Helical reconstruction
Fitted PDBs: 7x55
Q-score: 0.168
Koh A, Ali S, Popp D, Tanaka K, Kitaoku Y, Miyazaki N, Iwasaki K, Mitsuoka K, Robert R, Narita A
To Be Published
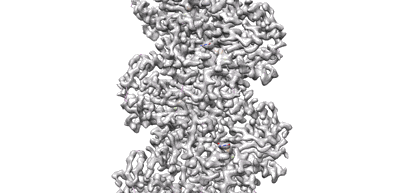
Resolution: 3.5 Å
EM Method: Helical reconstruction
Fitted PDBs: 7x56
Q-score: 0.486
Koh A, Ali S, Popp D, Tanaka K, Kitaoku Y, Miyazaki N, Iwasaki K, Mitsuoka K, Robert R, Narita A
To Be Published
- Magnesium ion (24 Da, Ligand)
- Guanosine-5'-diphosphate (443 Da, Ligand)
- Parm/stba family protein (32 kDa, Protein from Clostridium botulinum Bf)
- Cbg-parm filament with gdp (Complex from Clostridium botulinum)
A CBg-ParM filament with GTP or GDPPi
Resolution: 6.5 Å
EM Method: Helical reconstruction
Fitted PDBs: 7x59
Q-score: 0.189
Koh A, Ali S, Popp D, Tanaka K, Kitaoku Y, Miyazaki N, Iwasaki K, Mitsuoka K, Robert R, Narita A
To Be Published
- Guanosine-5'-triphosphate (523 Da, Ligand)
- Parm/stba family protein (32 kDa, Protein from Clostridium botulinum)
- Cbg-parm filament with gtp or gdppi (Complex from Clostridium botulinum)
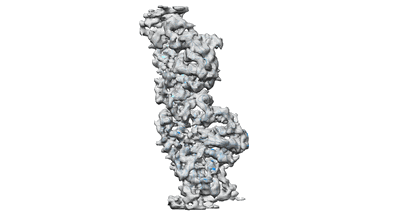
Whole structure of a 15-stranded ParM filament from Clostridium botulinum
Resolution: 4.7 Å
EM Method: Helical reconstruction
Fitted PDBs: 6izr
Q-score: 0.23
Koh F, Narita A, Lee LJ, Tanaka K, Tan YZ, Dandey VP, Popp D, Robinson RC
Nat Commun (2019) 10 pp. 2856-2856 [ Pubmed: 31253774 DOI: doi:10.1038/s41467-019-10779-9 ]
- Putative plasmid segregation protein parm (39 kDa, Protein from Clostridium botulinum Prevot 594)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Parm filament coded on pcbh plasmid from clostridium botulinum (Complex from Clostridium botulinum F str. 230613)
- Magnesium ion (24 Da, Ligand)
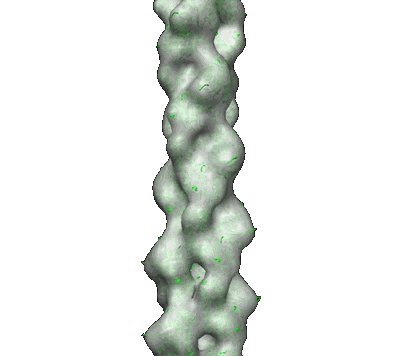
Averaged strand structure of a 15-stranded ParM filament from Clostridium botulinum
Resolution: 4.2 Å
EM Method: Helical reconstruction
Fitted PDBs: 6izv
Q-score: 0.362
Koh F, Narita A, Lee LJ, Tanaka K, Tan YZ, Dandey VP, Popp D, Robinson RC
Nat Commun (2019) 10 pp. 2856-2856 [ Pubmed: 31253774 DOI: doi:10.1038/s41467-019-10779-9 ]
- Putative plasmid segregation protein parm (39 kDa, Protein from Clostridium botulinum Prevot 594)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Parm filament coded on pcbh plasmid from clostridium botulinum (Complex from Clostridium botulinum F str. 230613)
- Magnesium ion (24 Da, Ligand)
actin-like ParM protein bound to ADP
Bharat TAM, Murshudov GN, Sachse C, Lowe J
Nature (2015) 523 pp. 106-110 [ DOI: doi:10.1038/nature14356 Pubmed: 25915019 ]
- Parm bound to adp (Sample)
- Parm (36 kDa, Protein from Escherichia coli)
actin-like ParM protein bound to AMPPNP
Resolution: 4.3 Å
EM Method: Helical reconstruction
Fitted PDBs: 5aey
Q-score: 0.398
Bharat TAM, Murshudov GN, Sachse C, Lowe J
Nature (2015) 523 pp. 106-110 [ DOI: doi:10.1038/nature14356 Pubmed: 25915019 ]
- Parm bound to amppnp (Sample)
- Parm (36 kDa, Protein from Escherichia coli)
CryoEM map of ParM filament at 7.2 Angstrom resolution
Resolution: 7.2 Å
EM Method: Helical reconstruction
Fitted PDBs: 4a6j
Q-score: 0.095
Gayathri P, Fujii T, Moller-Jensen J, van den Ent F, Namba K, Lowe J
Science (2012) 338 pp. 1334-1337 [ Pubmed: 23112295 DOI: doi:10.1126/science.1229091 ]
- Plasmid segregation protein parm (36 kDa, Protein from Escherichia coli)
- Parm filament (Sample)
Reconstruction of ParM in the closed state using cryo-EM
Galkin VE, Orlova A, Rivera C, Mullins RD, Egelman EH
Structure (2009) 17 pp. 1253-1264 [ DOI: doi:10.1016/j.str.2009.07.008 Pubmed: 19748346 ]
- Parm (Protein from unidentified)
- Parm (Sample)